By Alessandra De Cesare – University of Bologna
Microorganisms have key roles in both animal health and food safety. Their composition, level of diversity and temporal shifts in the animal’s gut and derived foods depend on many variables. We know that in the animal’s gut the microbial ecosystem is influenced by genetic parameters, rearing conditions, feeding strategies, use of therapeutic treatments and environmental conditions. We also know that the food microbial ecosystem is influenced by the food matrix, food processing and treatments as well as packaging and storage conditions. However, we miss precise information on the interactions between all those variables and microbes, meaning physiological and evolutionary data allowing robust predictions of microbial responses in specific conditions.
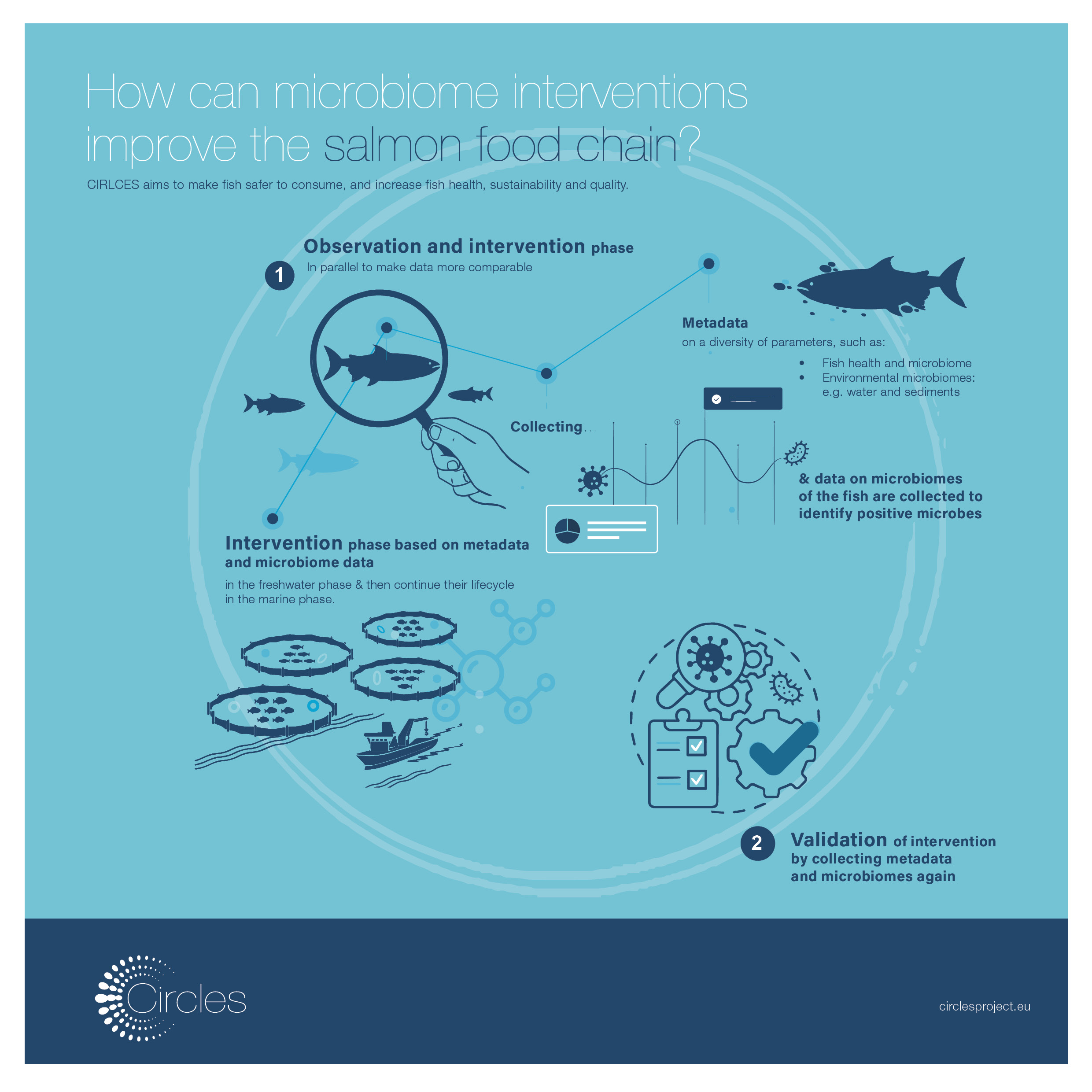
To understand and try to take advantage of positive microbe-based interactions, in CIRCLES we want to map and characterise the microbial ecosystems of selected food chains using microbiome research. In recent years, this field of research has witnessed a dramatic increase in scale and scope, due to the advances in sequencing technologies. Sequencing is becoming every day easier to access, thanks to the decreasing price and increasing quickness in obtaining results. When the human genome project started in 1990, using the Sanger sequencing, 13 years and 2.7 billion US dollars have been invested to sequence the human genome, which is about 3 billion base pairs. At present, by using the Next Generation Sequencing (NGS) technologies, 100.000 human genomes are roughly sequenced each year and the cost is less than 1000 US dollars/genome.
This “NGS revolution” has allowed to start many different international and interdisciplinary research programmes worldwide, aimed at the exploration of the microbial diversity in a large variety of ecosystems, including animals, foods and their related environments. To address studies involving microorganisms in animals and foods require an accurate profiling of microbial responses in representative conditions to be followed up by in-field observational tests in commercial situations. Indeed, we must improve our quantitative understanding of the animal and food microbiomes to identify the real drivers supporting animal productivity on one hand and food safety, quality and sustainability on the other hand.
In CIRCLES, labs in the field have been set up for the poultry and swine food chains to look at all the microbiomes circulating in those ecosystems; the animal microbiomes, the farm microbiomes, the farmer microbiomes and the food microbiomes. All these microbiomes are analysed using NGS sequencing in real samples collected at categorized times and conditions during the commercial rearing of the animals in real farms belonging to real poultry and swine companies. The aim is to understand which microorganisms and functions genes, for instance, the antimicrobial resistance genes, characterise those microbiomes, how they change on time or as a result of specific conditions, how they are influenced one by the other and what is the specific impact of each tested variable on the overall microbiomes circulating in the poultry and swine ecosystems. The interactions between microbiomes and all the variables we are registering are very difficult to establish. CIRCLES will overcome this problem by using good models and providing reliable predictions.
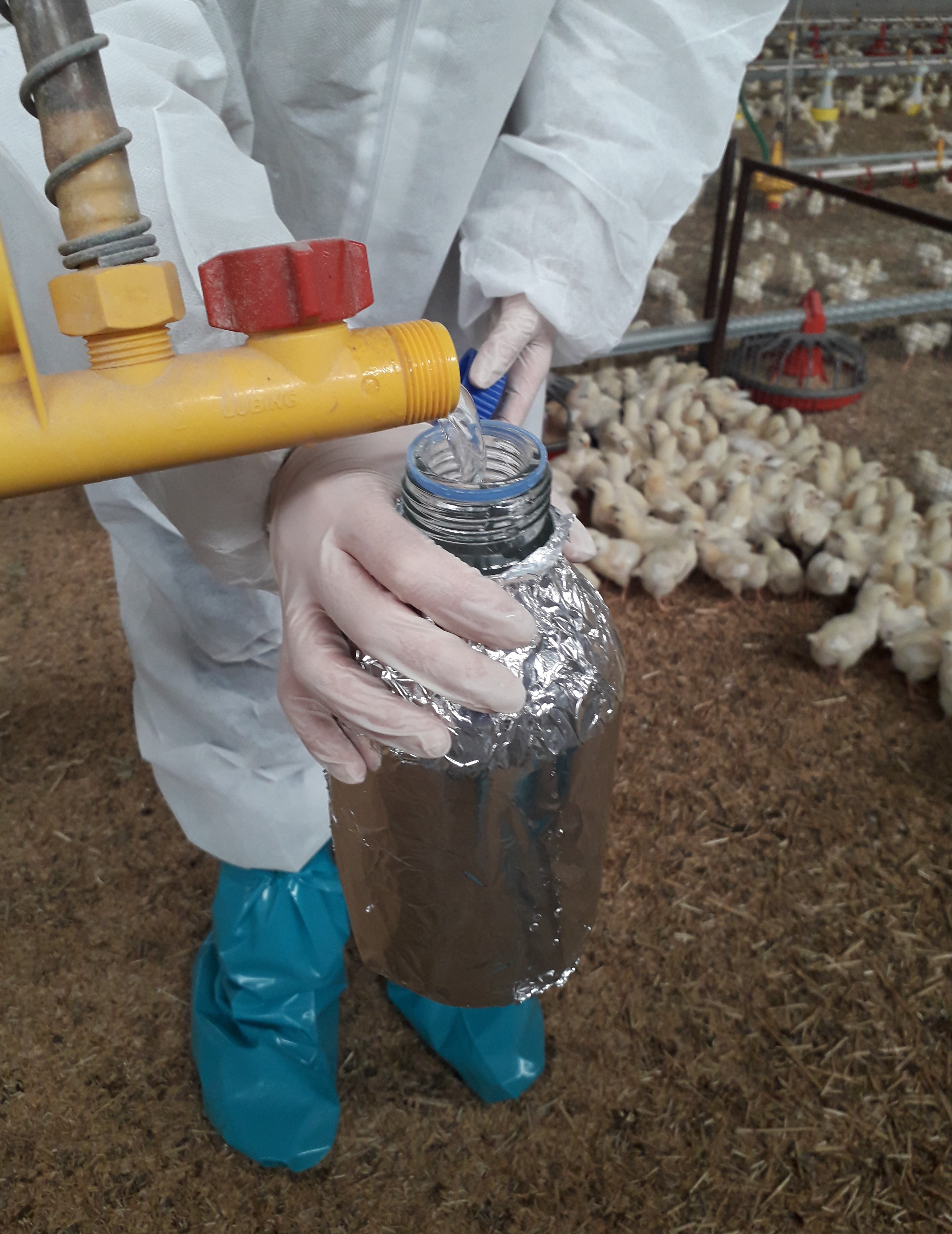
In the farms, we are sampling and sequencing animal faeces, feed and water, the air inside and outside the houses, soil outside the houses, faeces of the farmers and production wastes. At the slaughterhouse, we are sampling and sequencing meat products obtained from the animals investigated at the farm, food contact surfaces and environmental samples. The variables we are registering during samplings are the animal production parameters, temperature and relative humidity inside and outside the farm, feed conversion rate, animal body weight gain, animal mortality, use of whatever additive and antimicrobial for the animals or the litter, application of biosecurity measures, slaughtering time of the tested animals, cleaning procedures at the slaughterhouse, meat quality and microbiological parameters.
When all the results obtained by sequencing the animal microbiomes, the farm microbiomes, the farmer microbiomes, the food microbiomes and the microbiomes circulating in their surrounding environments will be available they will be analyzed to see which microorganisms and functional genes they contain. However, these results will be not considered as self-standing but always in connection with the variables registered when the poultry and swine samples have been collected by using quantitative models. The framework for quantitative models to correlated production and environmental variables to circulating microbiomes exists but to a large extent these models lack structured data on both animal and food microorganisms as those we are collecting in CIRCLES.
As results of our work of modeling sequences and categorized variables as well as groups of microbiomes and categorized variables we are sure to see trends, clusters, matchings coming out from our dataset to clarify on one hand which variables really affect keystone species or microbial communities supporting both animal health and food safety and, on the other hand, which variables can predict dysbiosis or some negative scenarios, such as infections in the animals requiring treatments or growth of foodborne pathogens in the meat.
We’ll make predictions based on data collected in real animals and meat products, sampled in commercial situations. This is not the end of the story because all our predictions will start to be validated back in the real poultry and swine meat companies in a couple of years. Hopefully, these predictions will be exploited at commercial level as new Smart Microbiome Food Products (SMFPs) with innovative Microbiome Transparent Labels (MTLs) for the poultry and swine food chains.
For centuries scientists tried to improve animal productivity, food quality and safety testing. CIRCLES will support improvements in productivity, quality, safety and sustainability of poultry and swine food chains by exploiting their beneficial microbes and positive interactions. Follow us to see how it ends!
Image credit: Dewdrop147 and Mutinka via Pixabay